But it may, if there is a good independent reason to think that X influences Y.
Correlation doesn’t imply Causation
Regression implies X causes Y
\[Y = slope \times X + intercept\]
- Both variables are continuous
- TONS of data are suitable for this kind of analysis.
- Examples?
Simple Linear Model
Simplest way two variables can be modeled as related to one another
\[Y_i = \beta_0 + \beta_1X_i + \epsilon_i\]
- \(\beta_0\) is the intercept (value of y where x= 0)
- \(\beta_1\) is the slope value expressing \(\Delta Y / \Delta X\)
- \(\epsilon_i\) is the error term (a normal random variable with mean 0 and variance \(\sigma^2\))
This equation should make sense
\[Y_i = \beta_0 + \beta_1X_i + \epsilon_i\]
Once you decide on a line, then the value of Y equals:
- the value predicted by the line, plus
- a random error from our error term
Finding the best line
- Many lines pass through \((\bar{X},\bar{Y})\). How do we pick the best one?
- There is unexplained variation in Y, so points don’t fall on straight line (why?)
Finding the best line - residuals
A residual represents the distance between the predicted value from the regression, and the actual value.
Another way to think about the residual: a single value from the normally distributed error term.
The squared residual is calculated as such:
\[d_i^2=(Y_i - \hat{Y}_i)^2\]
Finding the best line
The best line minimizes the residual sum of squares
\[ RSS=\sum\limits_{i=1}^n(Y_i - \hat{Y}_i)^2\]
We could do a Monte Carlo approach: try a bunch of slopes passing through \((\bar{X},\bar{Y})\) and calculate the RSS, then pick the smallest, but math offers a simpler solution.
Variances and Covariances
Recall the sum of squares: \[\ SS_Y = \sum\limits_{i=1}^n(Y_i - \bar{Y})^2\]
Sample variance: \[\ s^2_Y=\frac{\sum\limits_{i=1}^n(Y_i - \bar{Y})^2}{n-1}\]
SS equivalent to: \[\ SS_Y = \sum\limits_{i=1}^n(Y_i - \bar{Y})(Y_i - \bar{Y})\]
Variances and Covariances
With 2 variables, we can define sum of cross products \[\ SS_{XY} = \sum\limits_{i=1}^n(X_i - \bar{X})(Y_i - \bar{Y})\]
By analogy to the sample variance, we define sample covariance \[\ s_{XY} = \frac{\sum\limits_{i=1}^n(X_i - \bar{X})(Y_i - \bar{Y})}{(n-1)}\]
Sample Covariance - Negative
From \(-\infty\) to \(\infty\)
\[\ s_{XY} = \frac{\sum\limits_{i=1}^n(X_i - \bar{X})(Y_i - \bar{Y})}{(n-1)}\]
Sample Covariance - Positive
From \(-\infty\) to \(\infty\)
\[\ s_{XY} = \frac{\sum\limits_{i=1}^n(X_i - \bar{X})(Y_i - \bar{Y})}{(n-1)}\]
Calculate Covariance
By hand on whiteboard
x <- c(2 , 4, 3) y <- c(1, 5, 6)
Regression Parameters - Slope
In ordinary least squares regression (OLS), the slope of the best fit line is defined as: \[ \frac{covariance\ of\ XY}{variance\ of\ X}\]
Or: \[\hat{\beta}_1 = \frac{s_{XY}}{s^2_X} = \frac{SS_{XY}}{SS_X}\] note: this simplifies because both are divided by same denomenator
Regression Parameters - Intercept
The intercept is easy to find, because the best fit line passes through \((\bar{X},\bar{Y})\). Thus: \[ \bar{Y} = \hat{\beta}_0 + \hat{\beta}_1 \bar{X} \]
Or:
\[ \hat{\beta}_0 = \bar{Y} - \hat{\beta}_1 \bar{X}\]
Regression Parameters - Error
Recall: \[Y_i = \beta_0 + \beta_1X_i + \epsilon_i\]
Regression assumes that that \(\epsilon\) is a random normal variable with mean 0 and variance of \(\sigma^2\), which is related to the scatter around the line.
We estimate \(\sigma^2\) like this: \[\frac{RSS}{n-2}\]
The square root of this is standard error of regression: \[\hat{\sigma}^2=\sqrt{\frac{RSS}{n-2}}\]
Coefficient of Determination
In a linear relationship, the total variance we want to explain is \(SS_Y\).
Some variance is attributable to our error term (measured by \(RSS\)).
The remaining variance is explained by our regression, thus: \[SS_{reg}=SS_Y-RSS\]
Therefore: \[SS_Y=SS_{reg}+RSS\]
This is referred to as partitioning a sum of squares.
Coefficient of Determination
This leads to calculation of \(r^2\), AKA the coefficient of determination
\[r^2 = \frac{SS_{reg}}{SS_Y}=\frac{SS_{reg}}{SS_{reg} + RSS}\]
The square root of this value is known as \(r\) or the product-moment correlation coefficient.
Note: the sign of \(r\) comes from the sign of the slope of the line.
Hypothesis Testing
You will always get parameter estimates for the intercept and slope. The next question is: are they signficant?
The slope measures the strength of the effect of \(X\) on \(Y\). The slope is a measure of effect size
What is the null hypothesis about the slope in regression?
Hypothesis Testing ANOVA tables
| Source | Degrees of Freedom (df) | Sum of squares (SS) | Mean Square (MS) | F-ratio | P-value | |
|---|---|---|---|---|---|---|
| Regression | 1 | \(SS_{reg}=\sum\limits_{i=1}^n (\hat{Y}_i-\bar{Y})^2\) | \(\frac{SS_{reg}}{1}\) | \(\frac{SS_{reg}/1}{RSS/(n-2)}\) | F dist | |
| Residual | n-2 | \(RSS=\sum\limits_{i-1}^n(Y_i - \hat{Y}_i)^2\) | \(\frac{RSS}{(n-2)}\) |
- \(\bar{Y}\) = mean of Y
- \(\hat{Y}_i\) = predicted value from regression
- \(Y_i\) = value of Y
Confidence Intervals
We can also construct confidence intervals around our parameter estimates (formulae not shown).
Notice that our intervals are narrowest near the mean, and fatter the further you go away from mean.
Interpolation vs Extrapolation
Interpolation vs Extrapolation
Assumptions of Regression
- The causal relationship between X and Y is linear.
- The X variable is measured without error.
- Datasets that don’t meet this require Model II (a.k.a., RMA)
- Y values are independent with normally distributed errors
- Variance is constant along the regression line (homoscedasticity)
Doing Regression in R
myModel <- lm(response~predictor) summary(myModel)
## ## Call: ## lm(formula = response ~ predictor) ## ## Residuals: ## Min 1Q Median 3Q Max ## -6.332 -1.288 0.029 1.307 6.137 ## ## Coefficients: ## Estimate Std. Error t value Pr(>|t|) ## (Intercept) -1.082286 0.623277 -1.736 0.0828 . ## predictor 0.311143 0.006218 50.036 <2e-16 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 ## ## Residual standard error: 1.96 on 998 degrees of freedom ## Multiple R-squared: 0.715, Adjusted R-squared: 0.7147 ## F-statistic: 2504 on 1 and 998 DF, p-value: < 2.2e-16
Checking Assumptions: Residual plot
plot(lm(response~predictor))
Any systematic correlation between the residuals and the fitted is bad.
Checking Assumptions: Residual plot
Systematic departures from linearity can show up in the residual plots
response <- predictor^2 + rnorm(1000, sd=50) plot(lm(response~predictor))
Model II Regression - A different kind of residual
RMA regression assumes there is error in both X and Y
Minimizes an orthogonal residual sum of squares (along the major axis)
Show on white board
Doing Model II Regression in R
Model II regression requires a separate package lmodel2
In bioanthro literature, this regression is known as RMA (reduced major axis), but known in this package as MA (RMA in this package is something else)
The function is different, but the call is still simple.
Doing Model II Regression in R
library(lmodel2) results <- lmodel2(response~predictor) print(results)
## ## Model II regression ## ## Call: lmodel2(formula = response ~ predictor) ## ## n = 1000 r = 0.8318485 r-square = 0.691972 ## Parametric P-values: 2-tailed = 1.943335e-257 1-tailed = 9.716673e-258 ## Angle between the two OLS regression lines = 6.614198 degrees ## ## Regression results ## Method Intercept Slope Angle (degrees) P-perm (1-tailed) ## 1 OLS 0.8163301 0.2927504 16.31741 NA ## 2 MA -0.2438352 0.3033804 16.87677 NA ## 3 SMA -5.0856439 0.3519276 19.38838 NA ## ## Confidence intervals ## Method 2.5%-Intercept 97.5%-Intercept 2.5%-Slope 97.5%-Slope ## 1 OLS -0.3997441 2.032404 0.2806177 0.3048831 ## 2 MA -1.5022970 1.005865 0.2908500 0.3159985 ## 3 SMA -6.3165411 -3.896451 0.3400039 0.3642694 ## ## Eigenvalues: 108.3025 3.485197 ## ## H statistic used for computing C.I. of MA: 0.0001325621
In Class Breakout Group Exercise
20 minutes
Gordon et al. 2008

Gordon et al. 2008
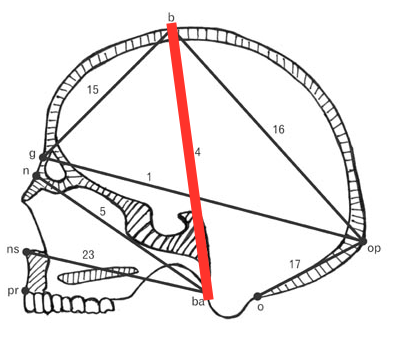
We are going to look at the scaling relationship between overall size and BBH (basion-bregma-height) in humans and fossil hominins
Exercise Step 1 - Import data
heads- Howell Craniometric Datafossils- Fossil Data
Exercise Step 2 - Calculate Geomean
Calculate the geomean for each individual in both data sets using the variables GOL, BBH, XCB, BNL, BPL, ASB
Add this data to a new column called geomean
geomean <- function(x, na.rm = TRUE) {
product <- prod(x, na.rm = TRUE)
n <- sum(!is.na(x))
geo <- product ^ (1/n)
return(geo)
}
Exercise Step 3 - Do Regression
Do an Ordinary Least Squares regression of log(BBH) as a function of log(geomean) for the modern data and a separate one for the fossils
Exercise Step 4 - Make Plots
Make a plot of log(BBH) as a function of log(geomean)
- plot the modern humans
- and add the modern human regression line to the plot
- add a new layer with the fossil data, being sure to differentiate them visually
- add the fossil regression line to the plot, using a different kind of line (e.g. dashed, or a different color)
Exercise Step 5 - Interpret
How do the fossils compare to the modern humans?
How does LB1 (H. floresiensis) compare to other species?